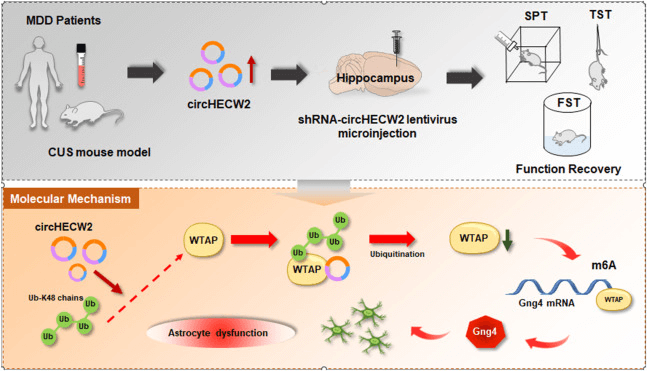
The research team led by Professor Yao Honghong from Southeast University has revealed the molecular mechanism by which CircHECW2 regulates the expression of WTAP and GNG4, thereby controlling the dysfunction of astrocytes.
Circular RNAs (circRNAs), a type of non-coding RNA, are generated by non-canonical back-splicing events and are highly expressed in the central nervous system (CNS). Numerous studies have shown that they are involved in many pathological and physiological processes.
In April 2024, the research team led by Professor Yao Honghong from Southeast University published an article titled "Engagement of N6-methyladenisine methylation of Gng4 mRNA in astrocyte dysfunction regulated by CircHECW2" in the journal Acta Pharmaceutica Sinica B (IF = 14.5). In this article, the authors found that the level of circular RNA HECW2 (circHECW2) was significantly increased in the plasma of patients with major depressive disorder (MDD) and in the mouse model of chronic unpredictable stress (CUS). Notably, the downregulation of circHECW2 could alleviate astrocyte dysfunction and CUS-induced depression-like behaviors. In addition, it was further demonstrated that the downregulation of circHECW2 increased the expression of the methyltransferase WTAP, leading to an increase in the expression of Gng4 through m6A modification, which provided functional insights into the correlation between circHECW2 and m6A methylation. This indicates that circHECW2 may represent a potential target for the treatment of MDD.
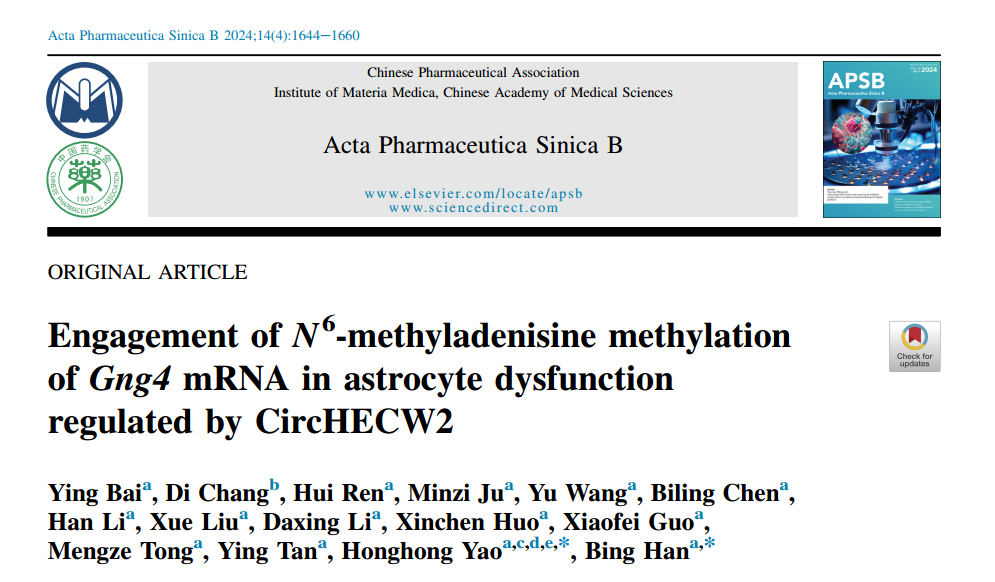

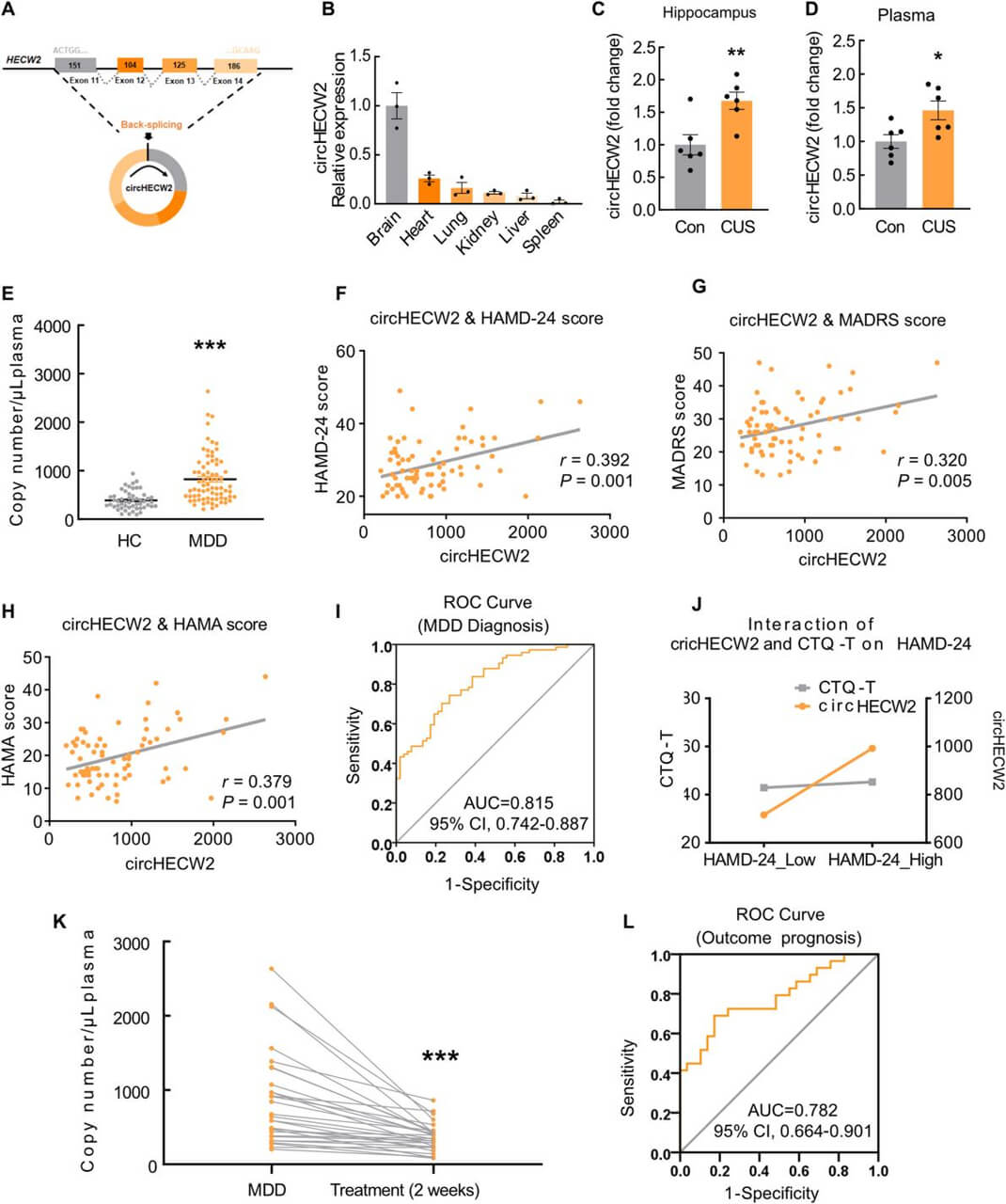
circHECW2 is upregulated in CUS mice and MDD patients
It is noteworthy that LPS- and CUS-induced models of depression are well-documented in studies relevant to depression. The author's previous study first demonstrated that circHECW2 levels were increased in the hippocampus of LPS treated mice. Thus, to investigate the potential involvement of circHECW2 in depression (Fig. 1C and D), the author isolated the hippocampus and collected plasma from CUS mice. Next, the author examined the levels of circHECW2 in the plasma of MDD patients and healthy control individuals (HCs), found that circHECW2 levels were markedly increased in MDD patients (Fig. 1E). Notably, our analysis revealed a positive correlation between circHECW2 levels and the scores of the Hamilton Rating Scale for depression 24 items (HAMD-24) (Fig. 1F), the scores of the Montgomery-Asberg Depression Rating Scale (MADRS) (Fig. 1G) and the scores of the Hamilton Anxiety Scale (HAMA) (Fig. 1H). Additionally, through linear regression analysis, the author uncovered that MDD patients with elevated circHECW2 expression levels and high scores on the childhood trauma questionnaire (CTQ) displayed more severe depression symptoms (Fig. 1J). To assess the predictive capacity of circHECW2 levels for MDD outcomes, the author examined the changes in circHECW2 level two weeks after treatment in MDD patient plasma and found that the level of circHECW2 was decreased 2 weeks after treatment in MDD patient plasma (Fig. 1K).
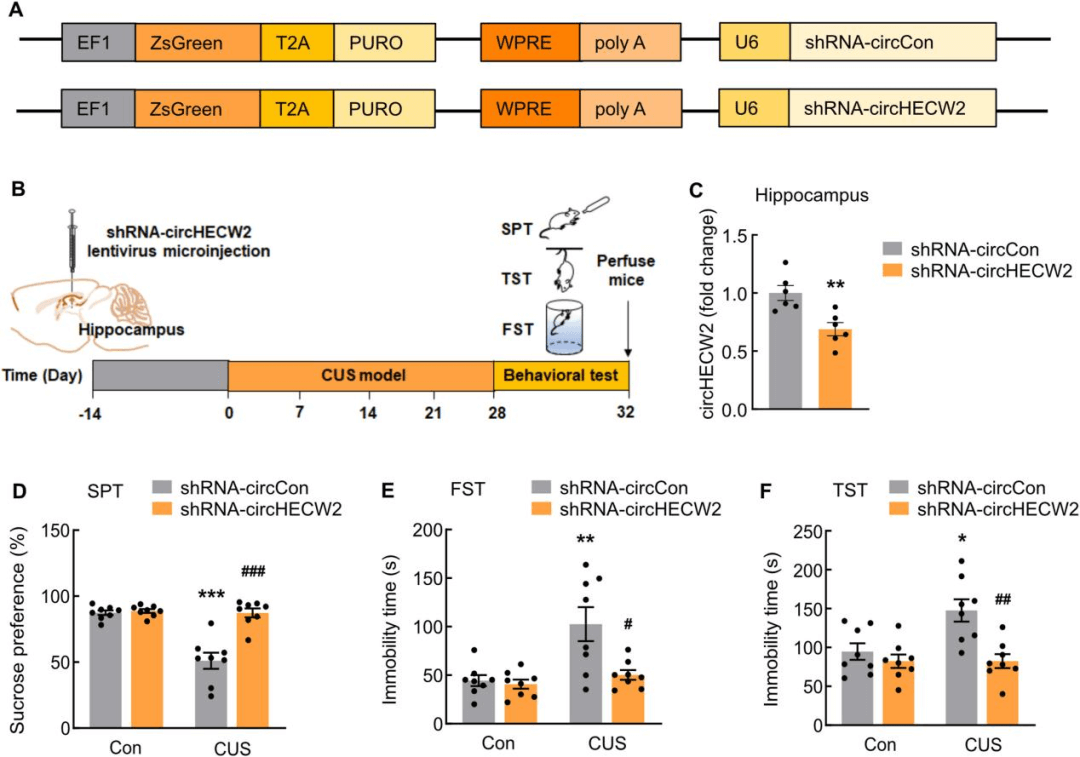
The downregulation of circHECW2 ameliorates the behaviors induced by CUS
After microinjection for 2 weeks, the author examined the efficacy of the lentiviral transduction and found that the expression of circHECW2 was decreased in shRNA-circHECW2-injected mice (Fig. 2C). Behavioural tests including sucrose preference test (SPT), forced swim test (FST), and tail suspension test (TST) were employed to evaluate the effect of circHECW2. The sucrose preference of the CUS-treated mice was decreased, indicative of anhedonia. Encouragingly, this deficit was significantly alleviated by the downregulation of circHECW2 expression (Fig. 2D). In both the FST and TST, the time of immobility was notably prolonged in the CUS mice, and these effects were markedly ameliorated in shRNA-circHECW2-injected mice (Fig. 2E and F).
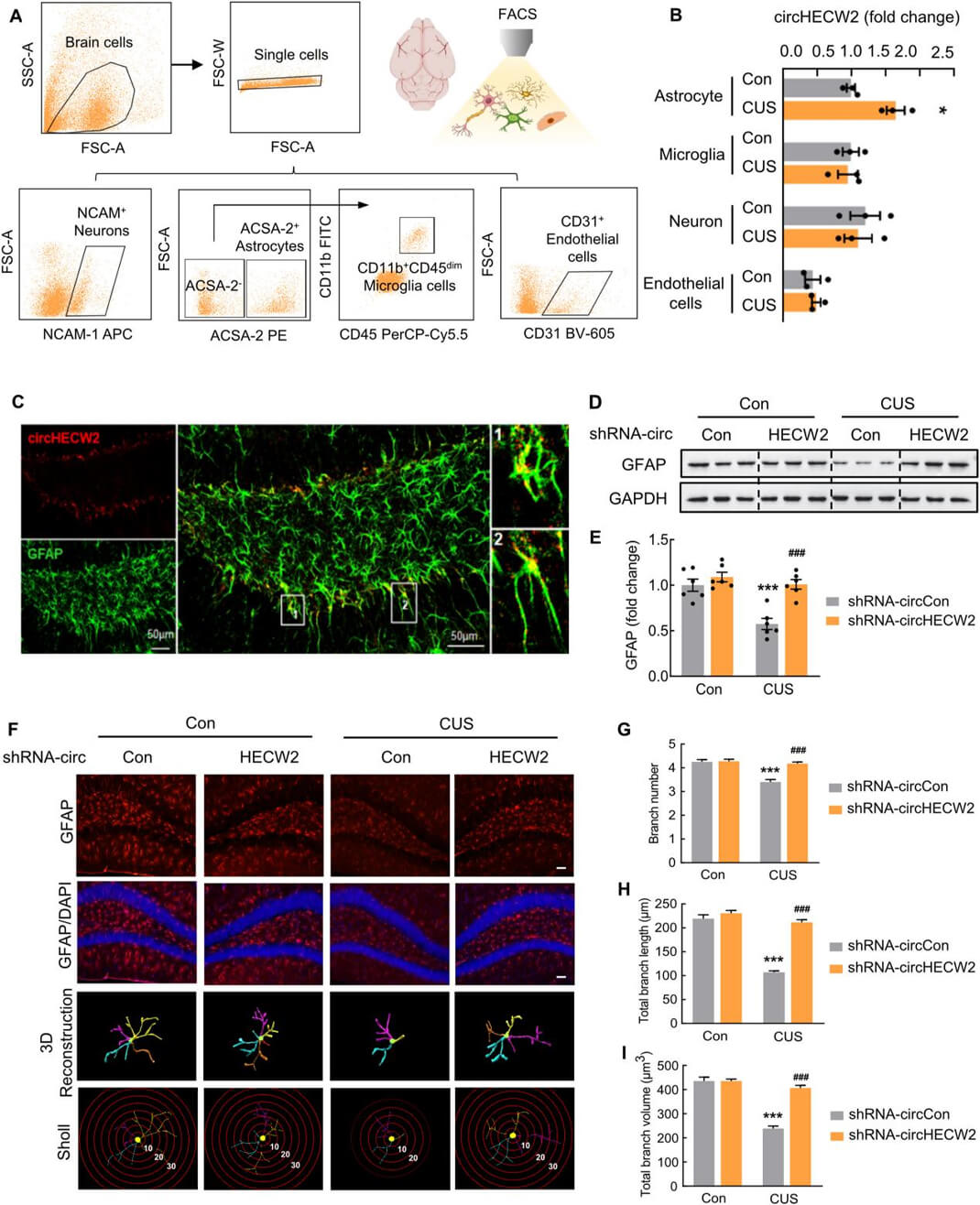
Role of circHECW2 on astrocyte dysfunction in CUS mice hippocampus
Subsequently, the author investigated the cellular mechanism through which circHECW2 affects functional recovery after CUS. To further assess the cell types in which circHECW2 expression is upregulated, the author detected the expression of circHECW2 in astrocytes, microglia, neurons, and endothelial cells from the CUS mice brain (Fig. 3A). The results revealed a significant upregulation of astrocyte-derived circHECW2 in CUS compared to microglia-, neuronal-, or endothelial cell-derived circHECW2 (Fig. 3B). Additionally, the fluorescence in situ hybridization staining indicated that circHECW2 was abundant in astrocytes (Fig. 3C). Furthermore, shRNA-circHECW2 treatment significantly mitigated the decrease in GFAP expression observed in CUS mice (Fig. 3D and E). Then, the author detected the function of shRNA-circHECW2 on the astrocyte’s morphology using GFAP and 3D reconstruction (Fig. 3F). Sholl analysis indicated that astrocyte dysfunction was induced by CUS, as evidenced by a reduction in branch numbers, length, and volume of astrocytes. Importantly, these deficits were markedly improved by shRNA-circHECW2 treatment (Fig. 3GeI). Taken together, these results suggest that the abnormal upregulation of circHECW2 in astrocytes may represent a critical molecular event in the progression of depression.
circHECW2 inhibits m6A methylation by downregulating WTAP
Given the potential role of m6A methylation in MDD and the mutual regulation between circRNAs and m6A methylation, we embarked on an investigation to determine whether circRNA’s regulatory role in the pathological processes of depression, particularly in astrocyte-mediated mechanisms, involves m6A modifications. CUS led to a decrease in m6A levels in the hippocampus, an effect that was significantly mitigated by shRNAcircHECW2, indicating a regulation of circHECW2 on m6A methylation (Fig. 4A). We next measured the expression of m6Amodifying enzymes above in CUS mouse models both in mRNA and protein levels, found that only the protein level of WTAP was reduced in the hippocampus of CUS mice (Fig. 4B). Subsequently, we used mouse primary astrocytes transduced with shRNA-circHECW2 lentivirus or circHECW2-overexpressed plasmid for further investigations(Fig. 4C and D). The expression of WTAP was significantly increased in shRNA-circHECW2-treated cells (Fig. 4E), whereas the Western blot analysis indicated that circHECW2 did not alter the levels of METTL3, METTL14, FTO, and ALKBH5, suggesting a specific association between circHECW2 and WTAP. To confirm these findings, primary astrocytes were transduced with the circHECW2 circHECW2-overexpressed plasmid and WTAP was significantly decreased in astrocytes (Fig. 4F). Next, pulldown assay was used to explore the interaction between circHECW2 and WTAP, and circHECW2 showed a stronger affinity to WTAP (Fig. 4G). Furthermore, in vivo experiments demonstrated that shRNA-circHECW2 significantly improved the decrease in WTAP expression induced by the CUS model (Fig. 4H). To explore why WTAP decreased after CUS, we employed immunoprecipitation to detect ubiquitination. The lysine 48-linked ubiquitination (Ub-K48) level of WTAP was significantly decreased by circHECW2 knockdown in the CUS model (Fig. 4I).
Next, we investigated the effect of circHECW2 and WTAP on the survival of astrocytes. Corticosterone was used to mimic the depression in vitro. Astrocytes transduced with shRNAcircHECW2 showed an amelioration of the decreased viability induced by corticosterone (Fig. 4J). In contrast, knockdown the expression of WTAP significantly aggravated the decreased viability of astrocytes treated with corticosterone (Fig. 4K). Furthermore, WTAP siRNA decreased viability of astrocytes was ameliorated by shRNA-circHECW2, further indicating a close relationship between circHECW2 and WTAP (Fig. 4L). Additionally, we constructed a brain astrocyte-specific AAVGFAP-WATP knockdown (KD) virus. Three weeks after the microinjection of AAV-GFAP-WATP KD and shRNAcircHECW2 lentivirus in the hippocampus, the mice were subjected to CUS or control. Behavioral experiments including SPT, FST, and TST were examined after 4 weeks of CUS exposure.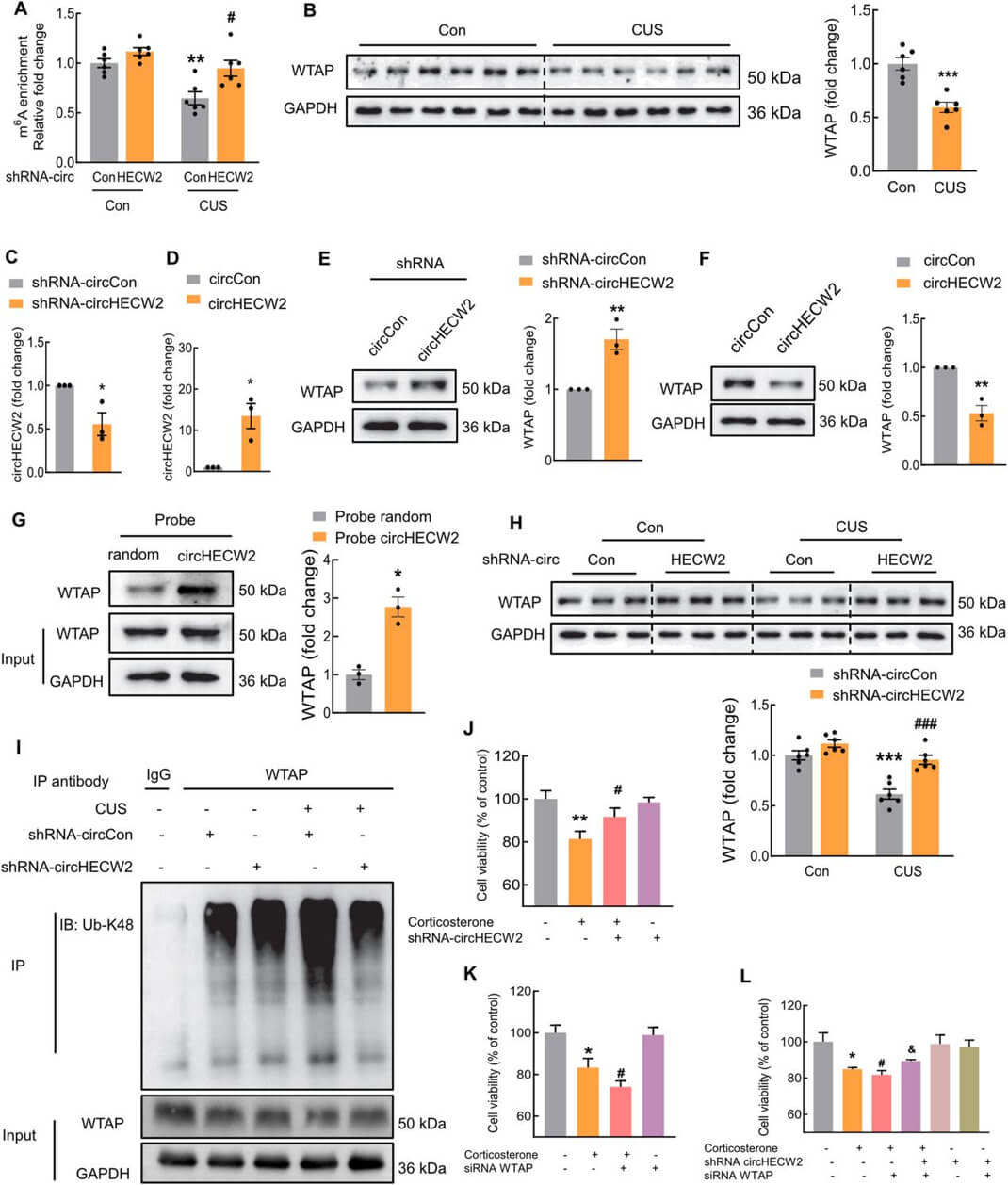
WTAP regulates m6A modification of Gng4 mRNA in depression
To seek the potential downstream molecule that participated in the CUS mouse, the author posted a transcriptome-wide detection of m6A modification in the hippocampus of the CUS mouse in previous study. The gene ontology biological processes (GO-BP) analysis showed that the downregulated genes were enriched in gene terms associated with the cellular process. The region of mRNA with altered m6A modification (downregulated genes) (Fig. 5B). Next, RNA-seq analysis was performed in the hippocampus of the CUS mouse model. We identified 288 genes that were differentially expressed in RNA-seq analysis (Cuffdiff adjusted P-value<0.05) (Fig. 5C). Of particular interest, there were 19 overlapping transcripts between the two comparisons (Fig. 5D), suggesting that these genes may be the target genes involved in astrocyte dysfunction after CUS. Notably, these 19 overlapping transcripts were verified at the mRNA level. In the 4 upregulated transcripts, no gene was validated. Additionally, GNG4 was significantly decreased in the plasma of MDD patients. Additionally, there was a negative correlation between GNG4 and the scores of the HAMD-Cognitive impairment. Based on these findings, GNG4 emerged as a central focus in our study of MDD. Furthermore, we analyzed the pathway involving GNG4 and its association with decreased m6A modification by Kyoto Encyclopedia of Genes and Genomes (KEGG) (Fig. 5E). Next, further results indicated that m6A modification was decreased in the peak region (chr13: 13825437e13825707) in the 30-UTR of Gng4 mRNA (Fig. 5F and G). The author employed luciferase assays to compare the wild-type (WT) and mutant m6A sites with and without WTAP siRNA treatment. These assays demonstrated that the mutation prevented methylation and increased the stability of Gng4 mRNA (Fig. 5H). As expected, overexpression of Gng4 markedly ameliorated the declined viability of astrocytes treated with corticosterone (Fig. 5I).
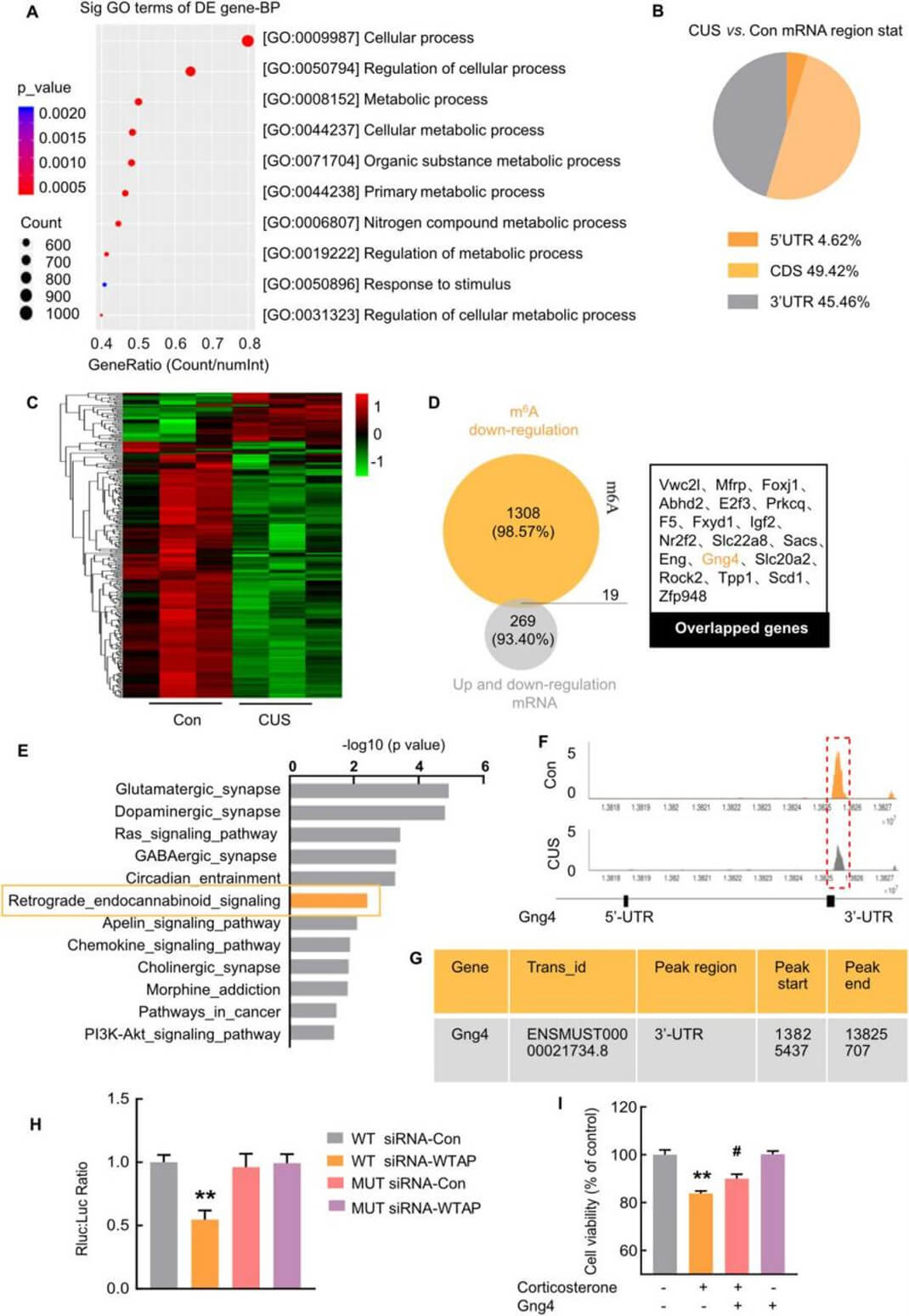
CircHECW2 regulates m6A methylation of Gng4 mRNA via WTAP
Furthermore, the author examined the expression and function of GNG4 in the CUS mouse model. The mRNA and protein levels of Gng4 were significantly decreased in the hippocampus of CUS mice, which was consistent with the decreased m6A modification of Gng4 (Fig. 6A and B). In addition, to understand the relationship between Gng4 levels and the WTAP, WTAP siRNA was transfected in mouse primary astrocytes. The expression of GNG4 and Gng4 mRNA are all decreased (Fig. 6C). Moreover, the upregulation of circHECW2 led to reduced mRNA and protein levels of Gng4 (Fig. 6D). Conversely, the downregulation of circHECW2 in astrocytes increased the mRNA and protein levels of Gng4 (Fig. 6E). Co-transfection of WTAP siRNA and shRNAcircHECW2 indicated that WTAP siRNA decreased the GNG4 level, which was markedly ameliorated by shRNA-circHECW2 (Fig. 6F). To validate these in vitro findings, Western blot analysis was performed to assess the levels of GNG4 in shRNA-circHECW2-treated CUS mice. The results showed that shRNA-circHECW2 treatment significantly mitigated the decrease in GNG4 expression observed in CUS mice (Fig. 6G and H). We also constructed a brain astrocyte-specific AAV-GFAP-Gng4 KD virus. Three weeks after the microinjection of AAV-GFAP-Gng4 KD and shRNA-circHECW2 lentivirus into the hippocampus, the mice were exposed to CUS or kept in a control condition. The results of SPT, FSF, and TST revealed that astrocytic Gng4 is involved in the regulation of circHECW2 in depression (Fig. 6IeK).
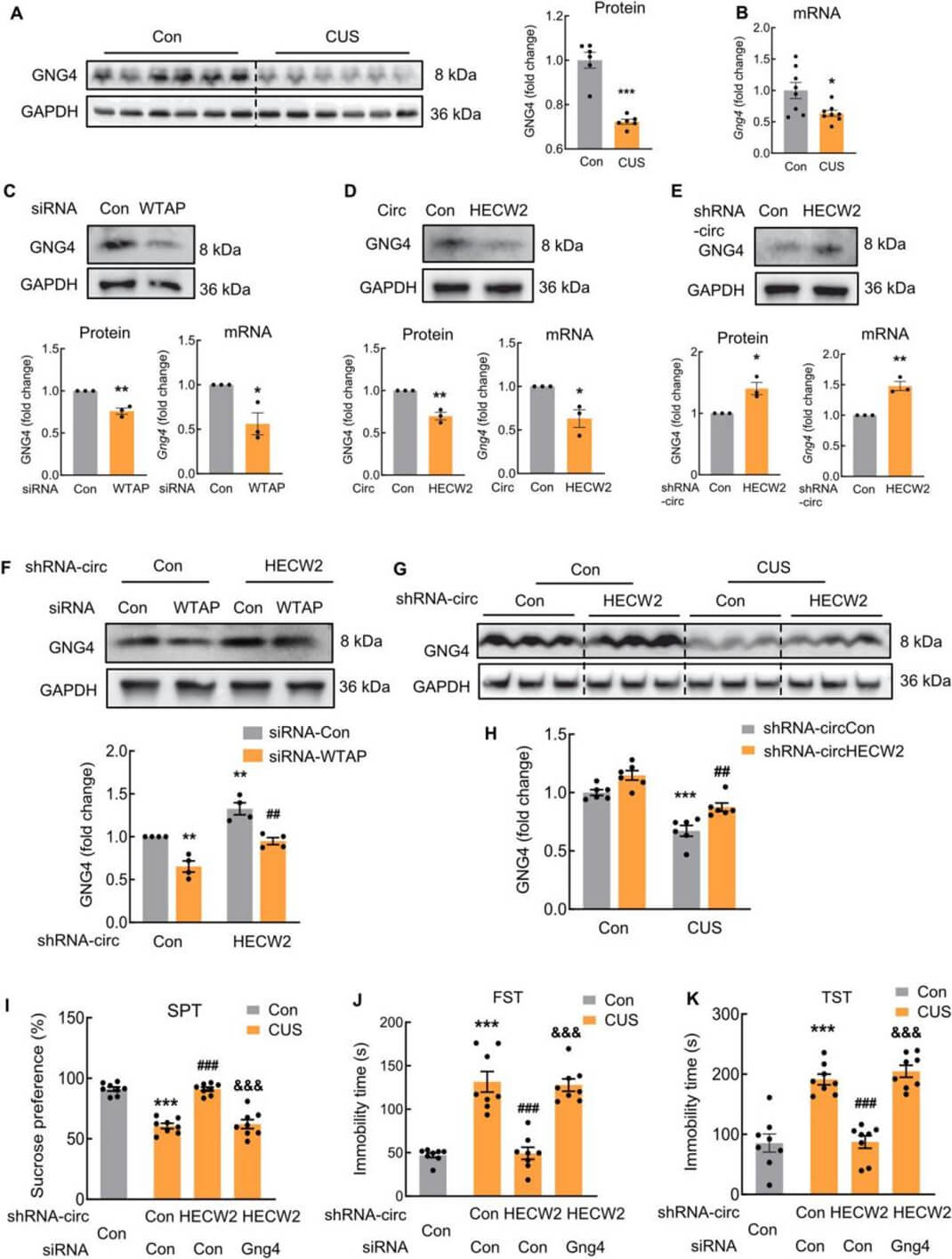
Hippocampus functional connectivity to the prefrontal cortex was positively correlated with GNG4 in MDD patients
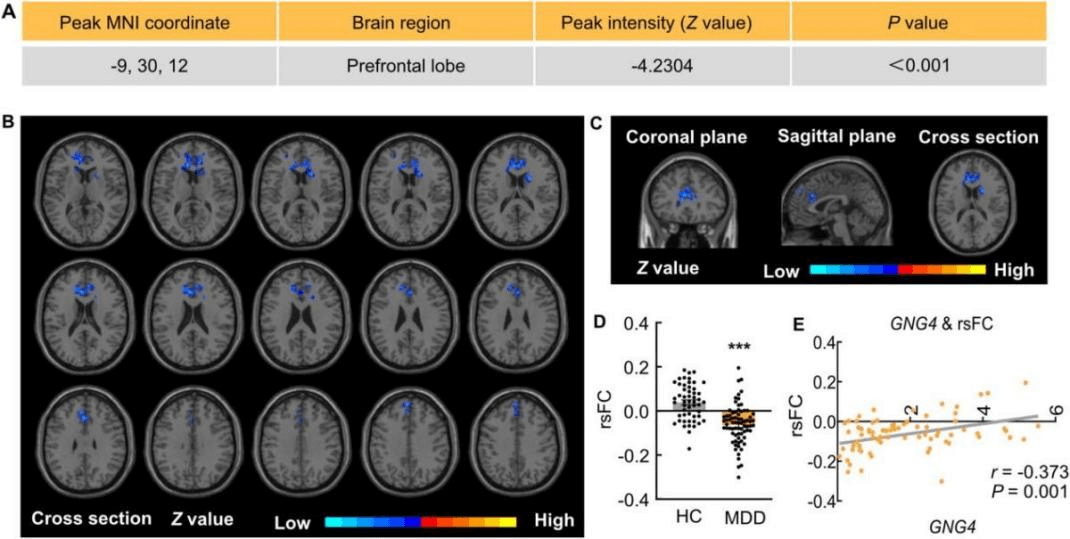
Finally, the seed-to-voxel analysis (hippocampal functional connectivity) was performed and showed hippocampus functional connectivity (FC) with prefrontal cortex (PFC) in MDD patients. As shown in Fig. 7A-C, the FC between the hippocampus and the prefrontal cortex was significantly reduced in MDD patients when compared to HCs. We next identified a noteworthy decrease in the resting-state functional connectivity (rsFC) between the hippocampus and the dorsolateral prefrontal cortex (Fig. 7D), whereas other brain regions, such as the visual cortex and inferior temporal lobe, exhibited no significant difference. Moreover, we found a positive correlation between GNG4 levels and rsFC, suggesting that GNG4 may play a role in the cognitive brain function of individuals with MDD (Fig. 7E).
Summary
Our study demonstrated that upregulation of circHECW2 led to the decrease in Gng4 mRNA via WTAP-mediated m6A modification, and caused subsequent astrocyte dysfunction. Specifically, circHECW2 promoted the ubiquitin-mediated degradation of WTAP in the CUS mouse model, and the effect of GNG4 on maintaining normal astrocyte functions is abolished when WTAP is downregulated in astrocytes. Therefore, the circHECW2/WTAP/GNG4 axis regulates astrocyte dysfunction by decreasing GNG4 stability via WTAP-mediated m6A modification. In conclusion, our findings indicate that circHECW2 holds promise as a therapeutic target for the treatment of depression. In addition, our study sheds light on the functional link between circHECW2 and m6A methylation, offering novel insights for the development of preventive strategies and effective treatments for MDD.